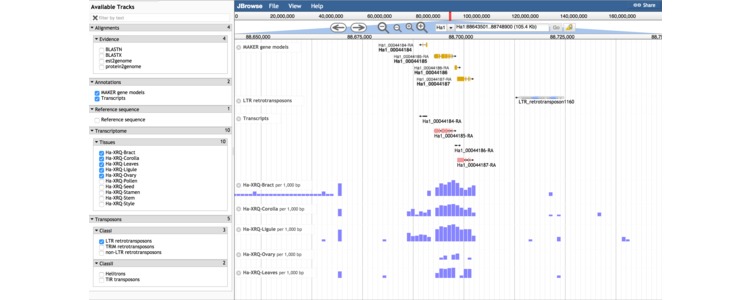
Shown above is a screen capture from the genome browser displaying expression levels of genes and transposons in a small window on chromosome 1.
Annotation of the sunflower genome has been a collaborative effort of researchers from INRA and UBC, and these efforts have resulted in the development of novel annotation software. Our approach, described briefly under the activities tab, relies on annotating and masking the repetitive fraction of the genome prior to annotating the gene space. The gene and transposon sequences, and the functional annotations, are available for download, viewing in the genome browser, and researchers may also perform a sequence-based search of the genome searching through the web-based BLAST tool which links results to the genome browser.
WARNING: This page is for reference. For all new projects, please use the latest version of the genome assembly and annotations.